Inferring delays in partially observed gene regulation processes
Hyukpyo Hong, Mark Jayson Cortez, Yu-Yu Cheng, Hang Joon Kim, Boseung Choi, Krešimir Josić, and Jae Kyoung Kim
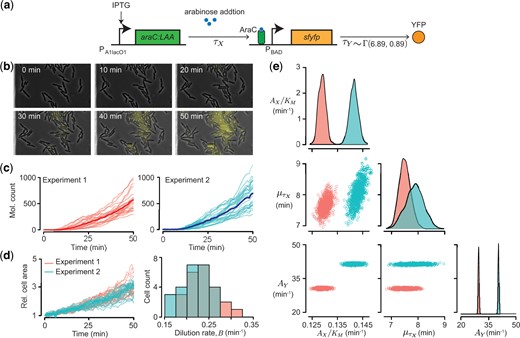
The authors proposed a simulation-based Monte Carlo Markov Chain method that accurately infers the parameters that describe the workings of a gene regulatory network, even when segments of these networks are unobserved. In their study, they demonstrate this method by examining a two-step process within cells: first, a signal activates the production of a hidden regulatory protein, which then triggers the production of observable fluorescent proteins. By leveraging knowledge about how these observable proteins are produced, their method allows them to understand how the hidden regulatory protein behaves over time. Through this approach, they were able to infer the time delay and various factors influencing the regulation of specific processes, such as the production of proteins from genes, even when these processes involve proteins that are not directly observable in experiments. Their method is adaptable and capable of handling complex models where certain components are hidden and cannot be directly observed.
See full paper here: https://doi.org/10.1093/bioinformatics/btad670
Complete Citation:Hyukpyo Hong, Mark Jayson Cortez, Yu-Yu Cheng, Hang Joon Kim, Boseung Choi, Krešimir Josić, Jae Kyoung Kim, Inferring delays in partially observed gene regulation processes, Bioinformatics, Volume 39, Issue 11, November 2023, btad670, https://doi.org/10.1093/bioinformatics/btad670